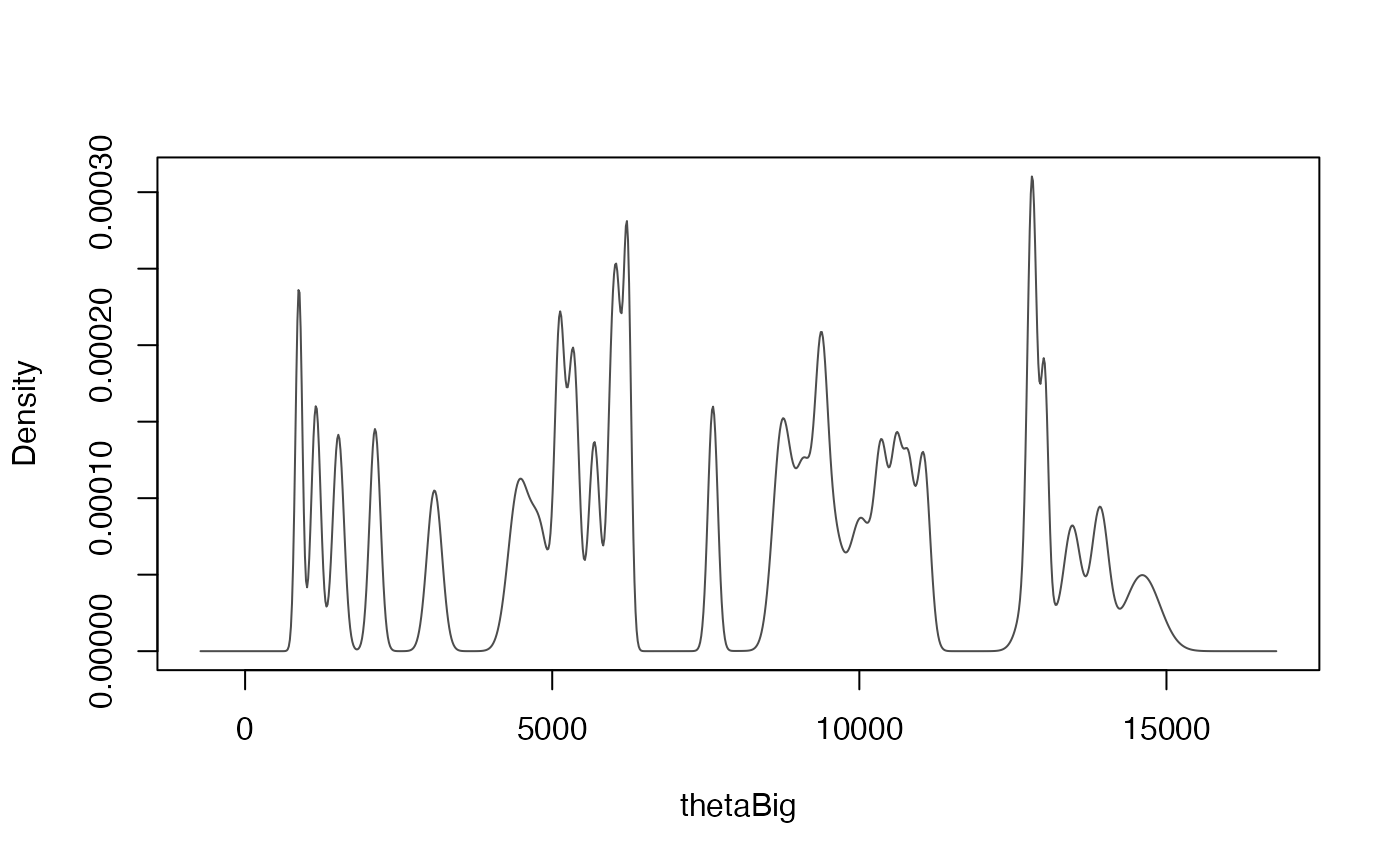
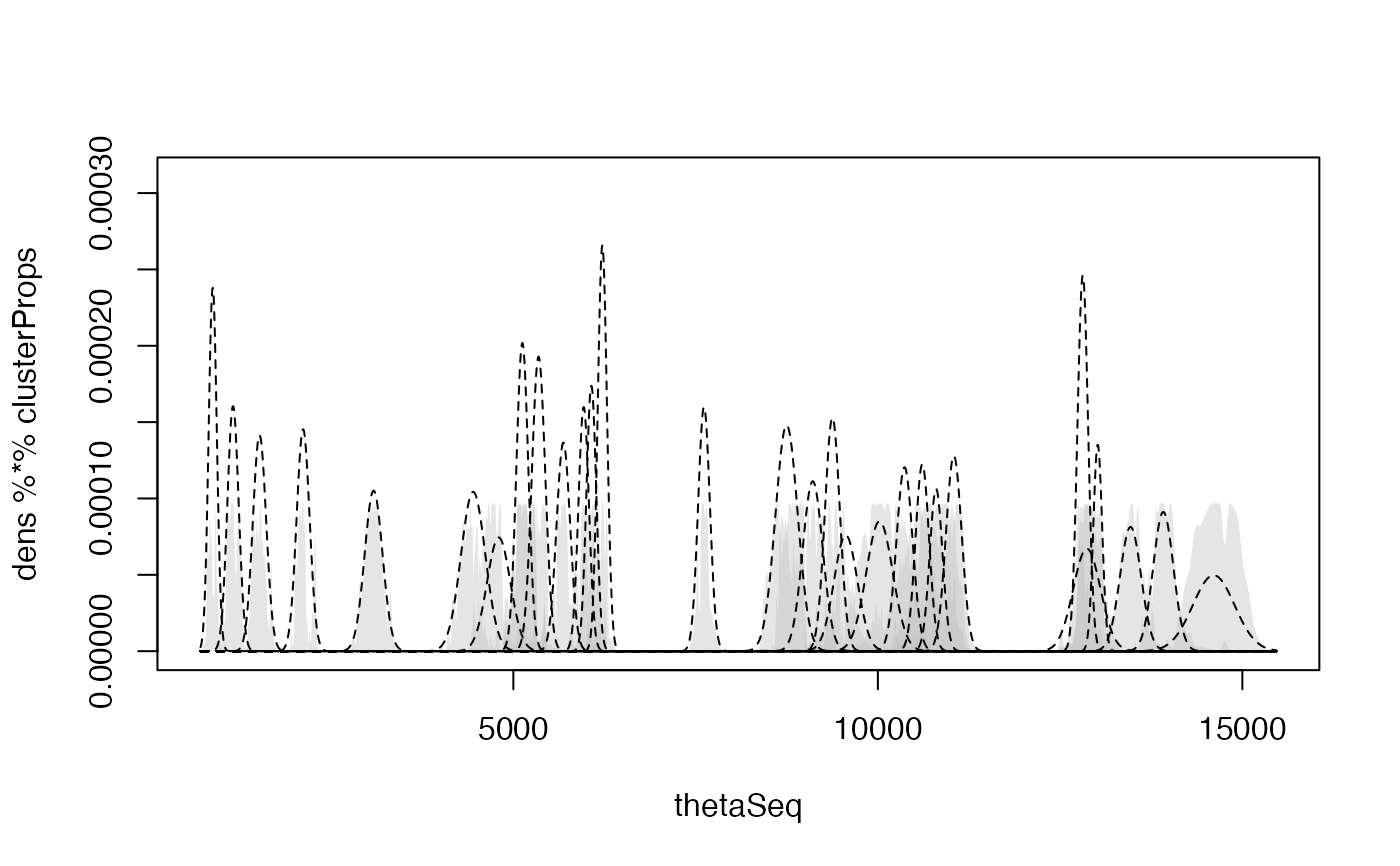
This function runs a non-parametric phase model on 14C and non-14C ages via Gaussian Mixture density estimation through the mclust package
BchronDensityFast(
ages,
ageSds,
calCurves,
pathToCalCurves = system.file("data", package = "Bchron"),
dfs = rep(100, length(ages)),
samples = 2000,
G = 30
)Arguments
- ages
A vector of ages (most likely 14C)
- ageSds
A vector of 1-sigma values for the ages given above
- calCurves
A vector of values containing either
intcal20,shcal20,marine20, ornormal(older calibration curves such as intcal13 are also supported). Should be the same length the number of ages supplied. Non-standard calibration curves can be used provided they are supplied in the same format as those previously mentioned and are placed in the same directory. Normal indicates a normally-distributed (non-14C) age.- pathToCalCurves
File path to where the calibration curves are located. Defaults to the system directory where the 3 standard calibration curves are stored.
- dfs
Degrees-of-freedom values for the t-distribution associated with the calibration calculation. A large value indicates Gaussian distributions assumed for the 14C ages
- samples
Number of samples of calibrated dates required
- G
Number of Gaussian mixture components
Value
An object of class BchronDensityRunFast with the following components:
- out
The output from the run of
densityMclustwith the given number of mixture components- calAges
The calibrated ages from the
BchronDensityfunction
Details
This is a faster approximate version of BchronDensity that uses the densityMclust function to compute the Gaussian mixtures for a set of calibrated ages. The method is an approximation as it does not fit a fully Bayesian model as BchronDensity does. It is designed to be a probabilistic version of the Oxcal SUM command which takes calibrated ages and sums the probability distributions with the aim of estimating activity through age as a proxy.
See also
Bchronology, BchronCalibrate, BchronRSL, BchronDensity for a slower exact version of this function